TIP-ChIP Service Overview
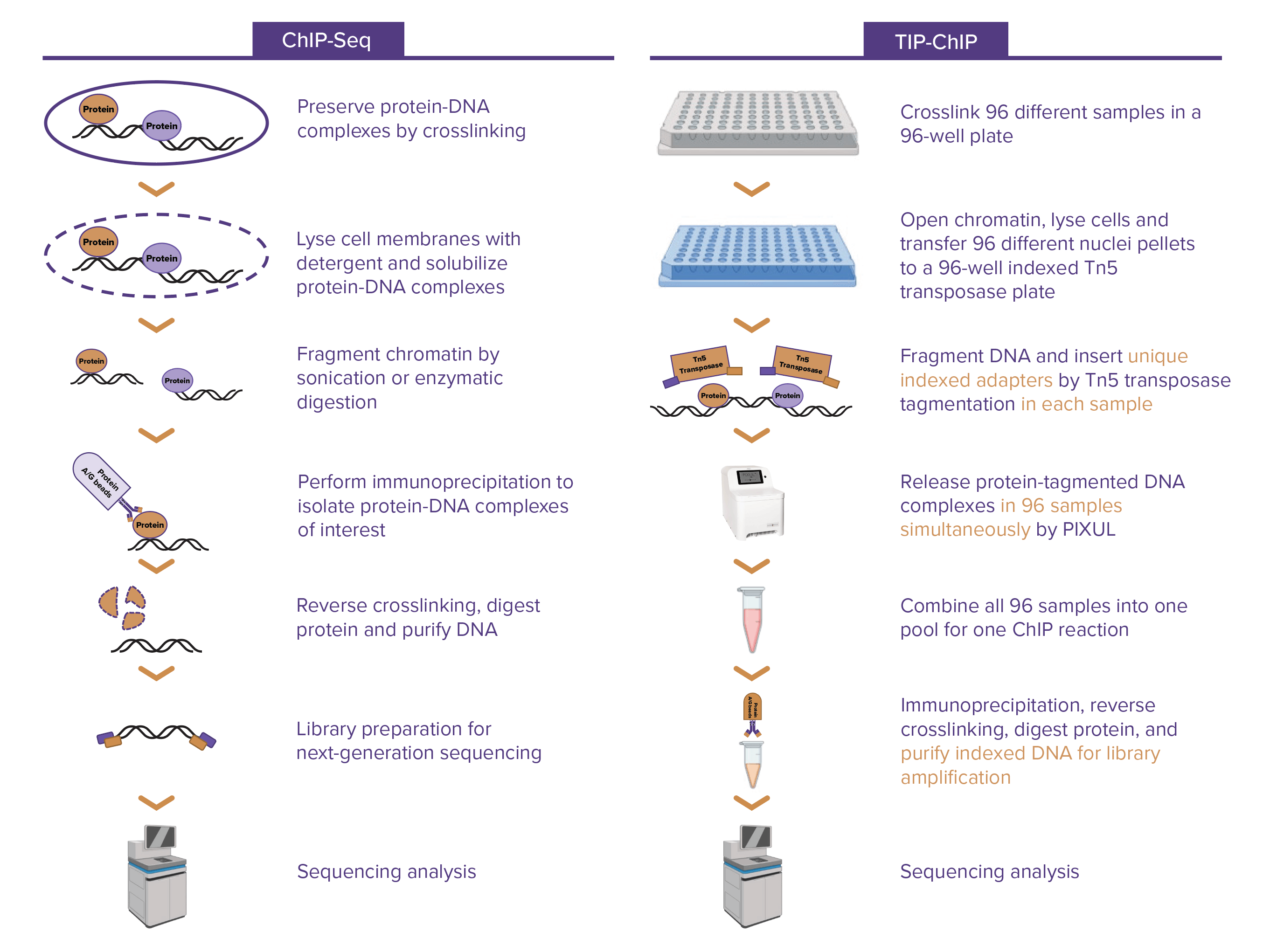
Tagmented, Indexed, and Pooled ChIP-Seq (TIP-ChIP) was developed for the rapid processing of 96 ChIP-Seq samples. The method is designed to reduce cell number requirements, increase throughput, and minimize sample-to-sample variability, offering a faster turnaround time (TAT), greater consistency across samples, and a lower cost.
TIP-ChIP starts with the unbiased opening and tagmentation of chromatin, inserting sequencing adapters across the genome in a 96-well plate format. The samples are then pooled into a single well, where the PIXUL® Multi-Sample Sonicator efficiently releases the the tagmented protein/DNA complexes. Next, the samples are transferred to a single tube for the immunoprecipitation reaction, seamlessly continuing the ChIP protocol to deliver high-quality, reliable results. We are excited to introduce our TIP-ChIP service with H3K27ac and H3K4me3 marks and we're continually expanding our portfolio of validated antibodies.
TIP-ChIP Highlights:
- Low cell input, recommended 300K to 1M cells per sample.
- Eliminate sample-to-sample technical variability
- High-Throughput format reduces batch-to batch variation
Comparison of TIP-ChIP and ChIP-Seq:
For experiments analyzing the same target across multiple samples, such as time-course or drug-response studies, or screening assays, TIP-ChIP provides a superior alternative to traditional ChIP. Its streamlined approach ensures consistent and reliable results across your sample set, eliminating sample-to-sample variability by processing all samples in a single ChIP reaction, making it the ideal choice for comparative analyses.
TIP-ChIP offers a distinct advantage with its low input requirement, needing just 300K to 1 million cells per sample, a fraction of the 1-10 million cells typically required for traditional ChIP. Additionally, its innovative tagmentation reaction assigns a unique identifier to each well of the 96-well plate. When samples targeting the same mark are pooled, only one set of sequencing primers is needed during PCR. These advantages enable the processing of up to 96 ChIP samples in just a matter of weeks, significantly accelerating your research timeline.
Like CUT&Tag and CUT&RUN, TIP-ChIP excels with low input material. However, TIP-ChIP stands out by delivering higher quality data at a more cost-effective price.
Active Motif’s End-to-End TIP-ChIP Service includes:
- Chromatin opening and cell permeabilization
- Tagmentation using Tn5 transposase for library generation
- PIXUL sonication
- Immunoprecipitation with a TIP-ChIP validated antibody
- PCR amplification
- Next-generation sequencing
- Comprehensive bioinformatics and data analysis
- Delivery of publication-ready figures
TIP-ChIP Service Data
Generation of TIP-ChIP Input Using Unbiased Tagmentation
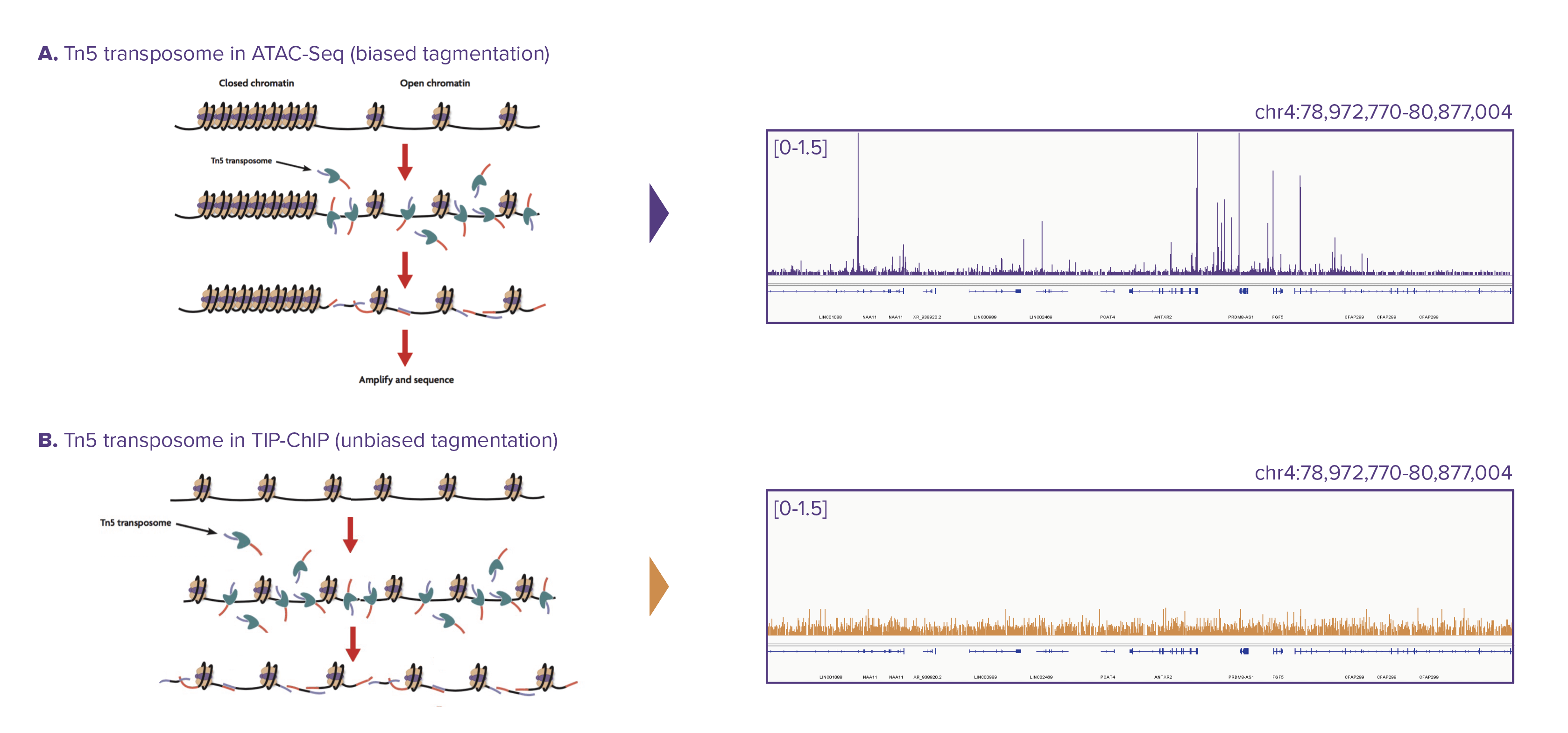
Figure 1: Tn5 transposomes used in biased and unbiased tagmentation.
Schematic of Tn5 tagmentation in A) ATAC-seq (biased) and B) TIP-ChIP (unbiased). In ATAC-seq, Tn5 transposomes can only access open chromatin, therefore the sequencing adapters are only inserted in those regions. Genome browser track generated using ATAC-seq is shown in purple where peaks are present only in the open chromatin regions. In TIP-ChIP, after cell fixation, the chromatin is all opened (secondary structure relieved) prior to tagmentation, resulting in sequencing adapters being inserted in an unbiased manner throughout the genome. Genomic browser tracks of TIP-ChIP input generated using unbiased tagmentation is shown in gold demonstrating even coverage across the genome.
Sample Multiplexing Using TIP-ChIP
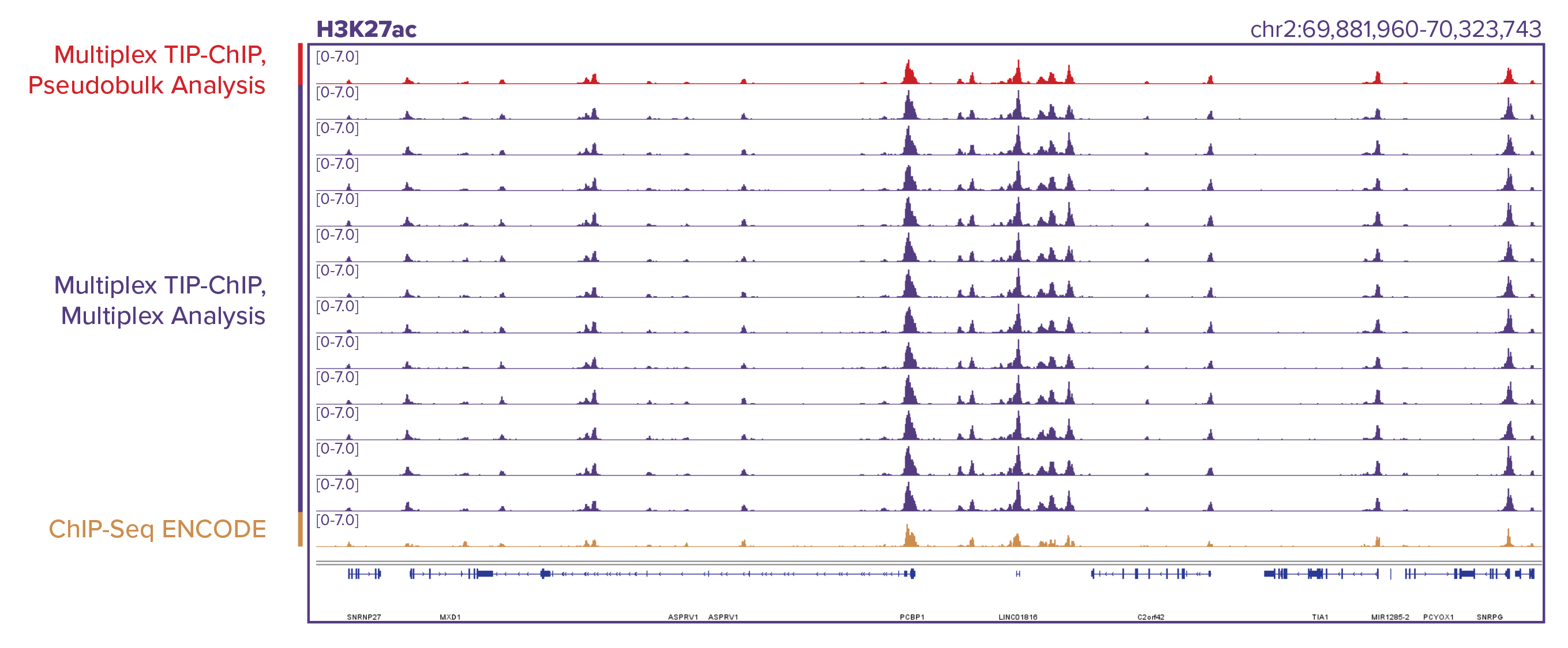
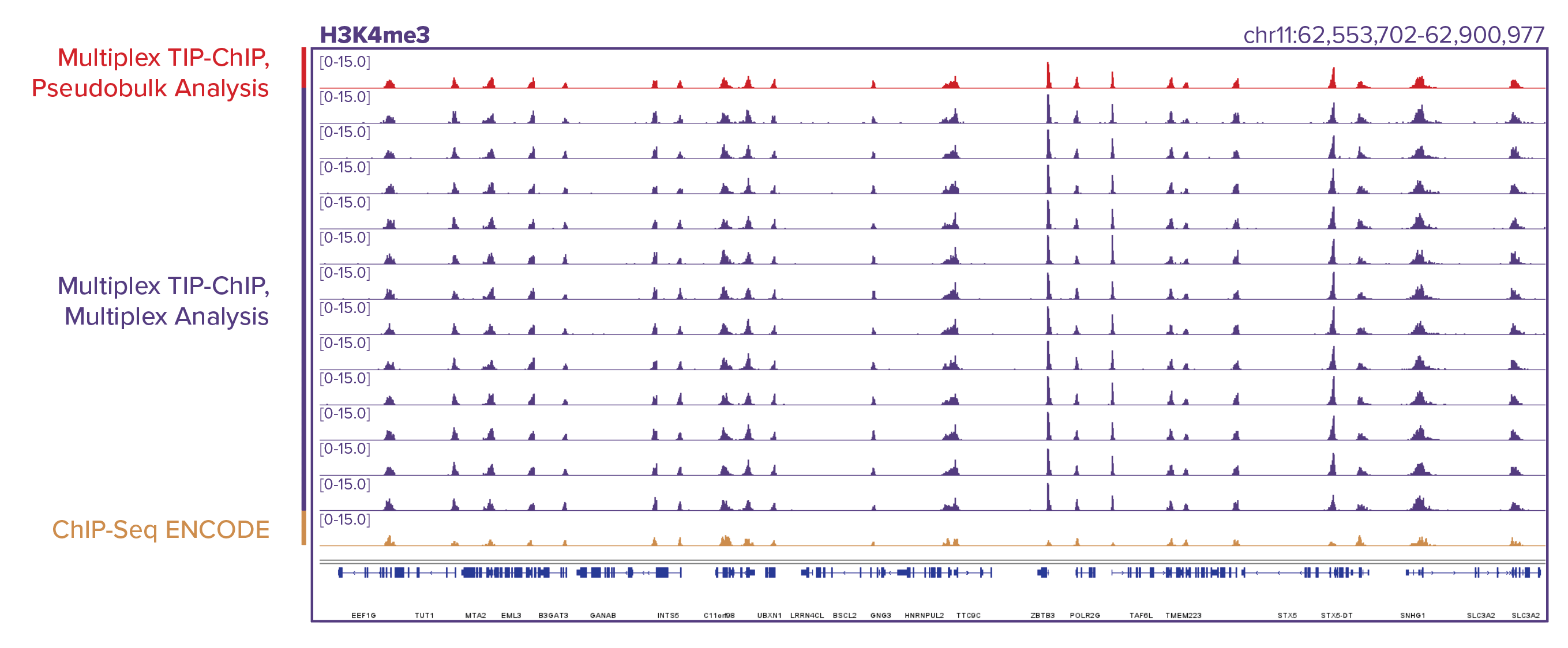
Figure 2: Comparison of TIP-ChIP and ChIP-Seq enrichment patterns.
Similar enrichment patterns are generated using TIP-ChIP and ChIP-Seq in the same cell line. TIP-ChIP was performed for both H3K27ac (top) and H3K4me3 (bottom) using 12 different K562 samples, ChIP-Seq data was taken from ENCODE. 12 individual samples were barcoded with Tn5, pooled into one ChIP reaction, and bioinformatically deconvoluted using TIP-ChIP barcodes. Deconvolution does not result in loss of resolution or data quality across the multiplexed samples. Genomic browser tracks were generated by first analyzing the 12 samples using a pseudo-bulk analysis method where sequencing data for all 12 samples is pooled without demultiplexing each uniquely barcoded sample (red tracks), and then by demultiplexing each of the 12 samples using a multiplex analysis method (purple tracks). The gold tracks are ENCODE genomic browser tracks generated using ChIP-Seq.
TIP-ChIP Reveals Differences in H3K27ac in PMA treated Neutrophils
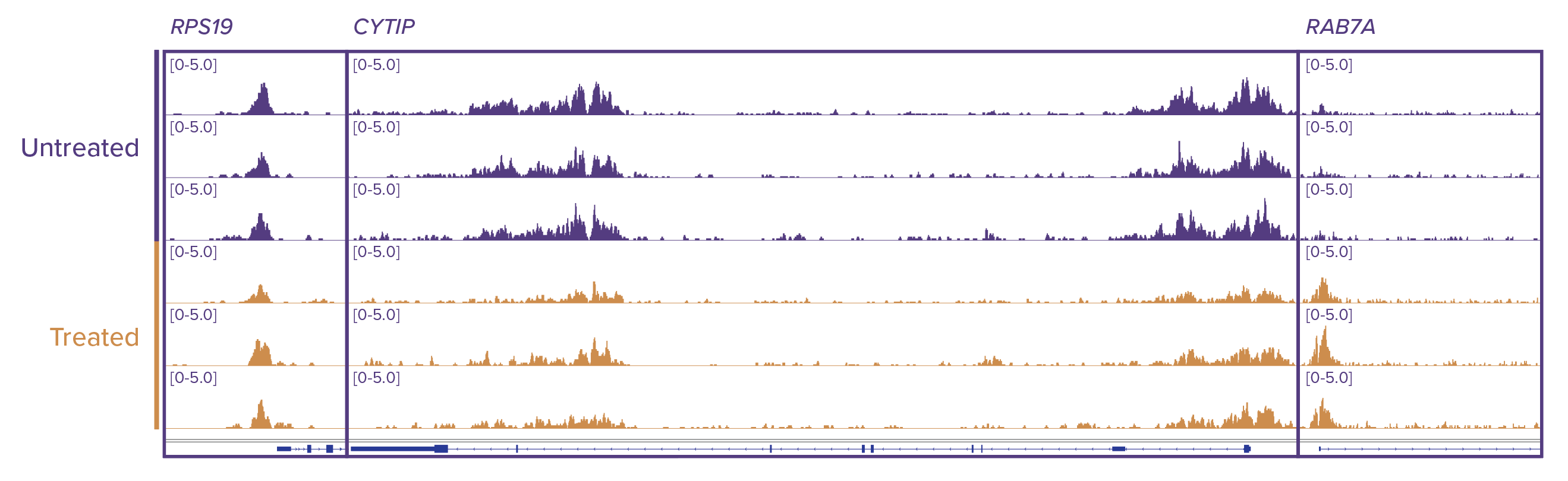
Figure 3: H3K27ac enrichment profiles at specific gene locations.
Genomic browser tracks showing enrichment patterns of H3K27ac at RPS19, CYTIP and RAB7A were generated using TIP-ChIP. Consistent local changes in H3K27ac enrichment are observed at CYTIP and RAB7A loci between treatment groups, but not at a housekeeping gene, RPS19. Purple tracks are neutrophils that were not treated with PMA, gold tracks are neutrophils treated with PMA for 60 minutes.
Sample Multiplexing Using TIP-ChIP
Figure 4: TIP-ChIP with H3K27ac can distinguish between treated vs untreated neutrophils.
A) Correlation plot of H3K27ac TIP-ChIP data generated using either untreated (T0) or 60-minute PMA treated (T1) neutrophils. Untreated samples 1-3 group together in the correlation plot while treated samples 4-6 group separately.
B) PCA analysis of H3K27ac TIP-ChIP data generated using either untreated (T0) or 60-minute PMA treated (T1) neutrophils shows clear separation of untreated and treated neutrophils.
TIP-ChIP Service FAQs
What are the sequencing details?
Is there a minimum sample number per reaction?
What is the minimum cell number per sample?
What is the most critical step in TIP-ChIP?
How can I determine if my antibody is suitable for TIP-ChIP?
TIP-ChIP Service Documents
TIP-ChIP Service Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.
You might also be interested in:
- View All Epigenetic Services
- [Blog] The latest ChIP Tip – How to Measure Single-cell Chromatin States with itChIP-Seq
- [Poster] TIP-ChIP: Tagmented, Indexed, and Pooled ChIP-Seq to Generate High-Throughput, Multi-Sample Results for Low Input Samples
References:
1. Ai, S., Xiong, H., Li, C.C. et al. Profiling chromatin states using single-cell itChIP-seq. Nat Cell Biol 21, 1164–1172 (2019). https://doi.org/10.1038/s41556-019-0383-5
2. Atteberry, B., et al., 2024. Front Immunol. 15. 144563
| Name | Cat No. | Price | |
|---|---|---|---|
| TIP-ChIP Service | 25211 | Get Quote |