Mod Spec® Service 概述
Mod Spec®是一种质谱分析方法,用于测定超过60种不同组蛋白修饰的相对丰度。Mod Spec®用于研究疾病响应和药物施用引起的表观变化是个不错的选择。检测得到的大量组蛋白修饰信息可帮助研究人员确定预期的表观变化,而且还能够帮助研究人员发现意想不到的变化,这可能会为研究提供更多有用且有趣的信息。
项目流程简述:
将收集好的细胞或组织交给我们Active Motif,我们将完成组蛋白提取、上机样品的准备、利用Thermo Scientific TSQ Quantum Ultra三重四极杆质谱联用仪进行样品的质谱分析。所有样本一式三份,数据以易于理解的格式呈现。在样本提交后的4-6周内我们将为您交付数据。
分析组蛋白修饰的变化:
- 对表观抑制剂的响应
- 正常 VS 病变细胞
- 敲除后的细胞或动物
- 抑制剂处理后的异种移植
- 人体活检样本
样本要求:
- 细胞量:2 – 5 百万细胞
- 组织量:25 – 100 mg
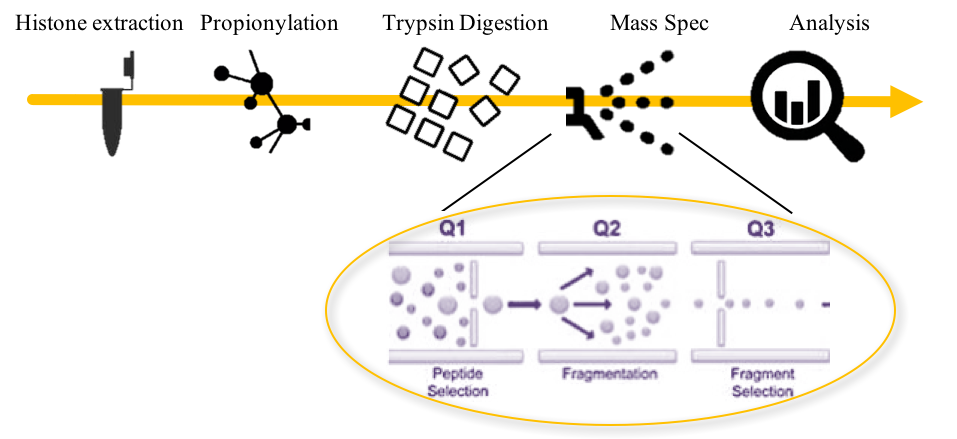
Mod Spec®️工作流程
1. 从样本中提取组蛋白
2. 用丙酸酐衍生化组蛋白。
- 蛋白质的质谱分析需要在注入质谱仪之前将蛋白质消化成短肽。然而,如果肽太小,它们不能在LC步骤中分离,如果太大,它们太复杂,无法分析。最常见的消化方法是胰蛋白酶,这是一种在赖氨酸和精氨酸残基处分解蛋白质的酶。但这在分析组蛋白时是一个问题,因为组蛋白尾部富含赖氨酸,导致消化的肽太小。使用丙酸酐的丙酰化可以阻止赖氨酸的裂解,同时允许精氨酸的消化。所得肽的大小适合质谱分析。
3. 组蛋白被胰蛋白酶消化。
4. 样品在三重四极杆质谱仪上进行分析。
- 筛选适当分子量的肽。
- 肽在碰撞室中进一步碎裂
- 选择感兴趣的肽片段并通过质谱分析
5. 对碎片进行分析并绘制图表,以显示样本之间的变化。
Histone Modifications Detected by Mod Spec®
^ 多种H2A亚型可能有贡献了H2A修饰的信号
* H3R2me2和H3K4me2/3是相互排斥的修饰。H3R2甲基化阻止H3K4甲基化。因此,H3K4修饰仅在H3R2未修饰肽上呈现。有关更多信息,请参阅 Nature. 2007 Oct 18; 449(7164):933-7.
♦ H3.2可能会显示出H3.1的信号
客户对我们服务的评价:
"在研究表观遗传药物的分子机制时,我们将Mod Spec®外包给了Active Motif,以便更广泛地了解药物诱导的组蛋白修饰变化。总的来说,我们对服务的质量、项目周期以及价格非常满意。我们发现了我们以前没有注意到的令人惊奇的结果。"
Matthias Lauth, PhD
菲利普斯大学
马堡,德国
查看完整的推荐信列表 >
Mod Spec®️服务数据示例
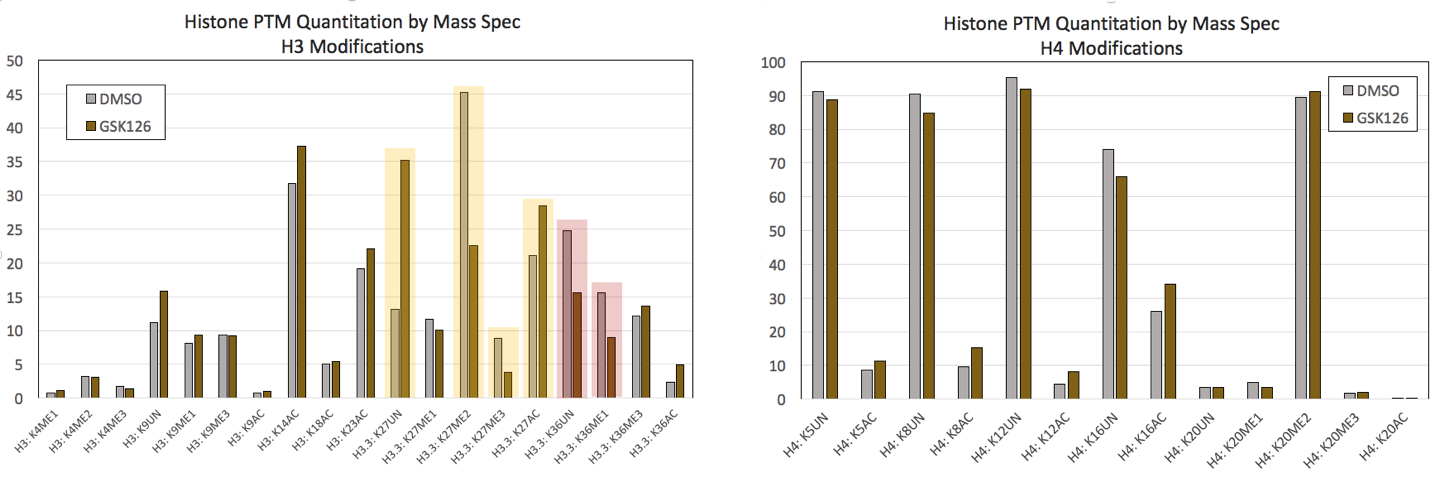
图1:来自对照HeLa细胞和用0.5 μM GSK-126处理7天的HeLa细胞的Mod Spec®数据。(点击图片可放大)
GSK126是一种阻断EZH2甲基转移酶活性的抑制剂,导致H3K27me2和H3K27me3的整体降低。A) 从Active Motif的Mod Spec®分析中可以选择H3修饰,确认H3K27甲基化显著降低,同时乙酰化和未修饰的H3K27(以黄色突出显示)增加。还检测到H3K36的变化(以红色突出显示)。B) 选择的H4修饰对GSK126处理的反应没有显示出显著的变化。
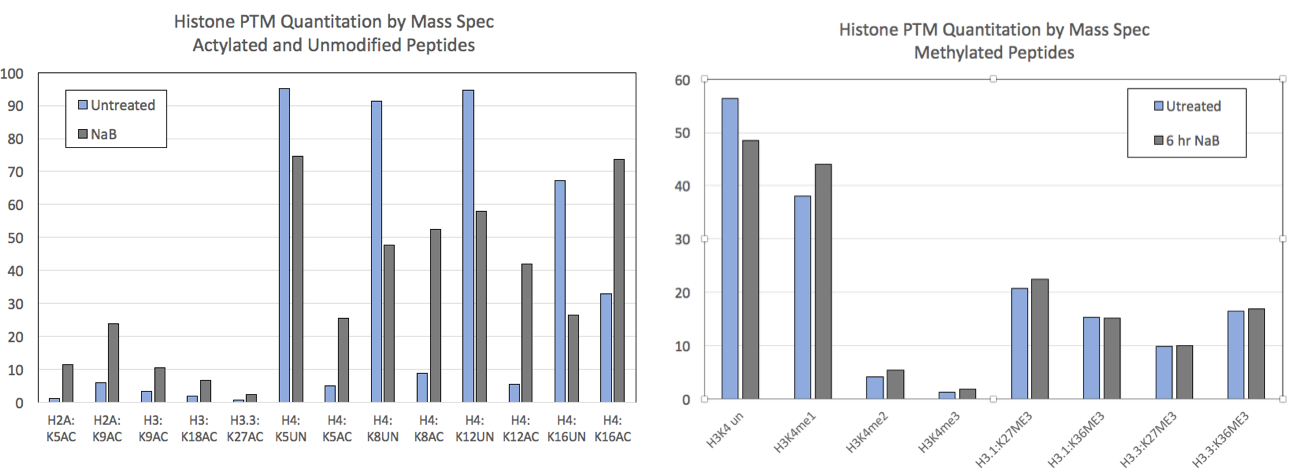
图2:Mod Spec®数据来自对照HEK293细胞和用5 mM丁酸钠(NaB)处理6小时的HEK293细胞。
NaB是一种通用的HDAC抑制剂,NaB处理可提高组蛋白乙酰化水平。左图 显示了选择乙酰化和未修饰组蛋白位点的Mod Spec®数据。处理后所有位点的乙酰化都增加。未经修饰的肽在某些位置有信号,这些位置并在NaB处理后显示相应的减少。B) 不同组蛋白残基的甲基化不受NaB处理的影响。
Mod Spec®️服务文献
搜索我们的数据库中使用了我们Mod Spec®服务的用户的文献。
Mod Spec® Service 文件
Mod Spec® Service Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.
You might also be interested in:
| Name | Cat No. | Price | |
|---|---|---|---|
| Mod Spec® | 25085 | Get Quote | |